Karnataka 2nd PUC Biology Notes Chapter 11 Biotechnology: Principles and Processes
→ Biotechnology is a synthetic co-integrated branch of science which utilises the knowledge of biochemistry, chemical engineering and microbes, in tapping the inactive behaviour of microbes and cultured cells for technical applications.
→ It can be defined as a technology in which microorganisms and isolated cells, tissues and organs of desired plants are employed to produce novel varieties of desired products for the welfare of mankind. The term biotechnology was coined by “Karl Erecky”.
![]()
Principles of Biotechnology:
I. Genetic engineering or Genetic surgery:
The method of isolating or synthesizing the desired gene and its insertion in the genome of the other organisms to produce hybrid DNA or recombinant DNA is called genetic engineering.
II. Maintenance of sterile environment
Tools used in genetic engineering are as follows.
- Desired gene
- Vector or Vehicle DNA
- Enzymes.
- Host cell
- Bioreactors.
1. Desired gene: The gene of interest which is also called desired gene or foreign DNA, has to be obtained from its source or is to be synthesized artificially in the lab.
2. Vectors: These are molecular vehicles which carry the desired gene from one organism to the other organism. There are different types of vectors namely plasmids, bacteriophages, cosmids, phasmids etc [cosmid means plasmid + cohesive ends of phage DNA, Pasmid means phage DNA+a small part of plasmid]
Plasmids:
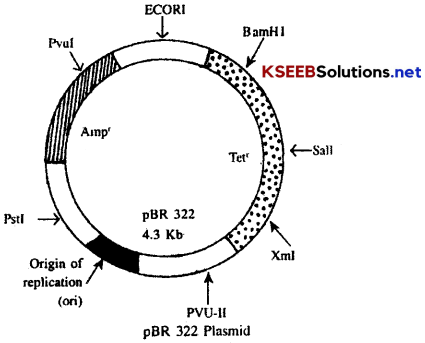
The small circular and extra chromosomal double stranded DNA molecules present in the cytoplasm of bacteria are called plasmids. They show self replication. They contain 2 to 8 thousands of nitrogen base pairs. They also contain some genes, like antibiotic resistant genes and genes for expression of some characters. They can be cut at specific sites using restriction enzymes and desired genes can be inserted into these plasmids. There are different kinds of plasmids, e.g. F-plasmid [Sex, plasmid], R-plasmid [resistant plasmid], col plasmid [colicins-that kill some strains]. In additions to these there are some genetically engineered plasmids like pBR-322 [Plasmid of Boliver and Rodriguez] and pUC-18 [Plasmid of University of California]
![]()
Structure of pBR-322:
pBR-322 is a natural plasmid and F+ plasmid. It is about 4.3 kb in size. It is a plasmid with an ori site (origin of replication), two antibiotic resistance sites, selectable markers for Amphicillin resistance (Ampr) and Tetracycline resistance (Tetr). It has thirteen unique sites in different regions out of which seven are important. A unique site is a specific restriction enzyme (REN) recognition site called ECORI.
Vectors for cloning genes in plants and animals are Ti plasmid isolated from Agrobacterium tumefaciens in plants and retrovirus are now made non pathogenic and are used to deliver gene into animal cells.
Note: Insertional inactivation: When a gene or recombinant DNA is inserted within the coding sequence of a vector, the coding sequence responsible for an enzyme or a particular character becomes inactivated. This is known as Insertional inactivation.
3. Enzymes: There are two main types of enzymes that are mainly used in the genetic engineering. They are RENs and Lygases.
(a) Restriction endonucleases-REN: The nuclear enzymes that break DNA at specific sites are called restriction endonucleases [REN], The RENs are popularly known as Biomolecular scissors. These are first discovered by Hamilton smith. There are a number of restriction endonucleases. Each enzyme recognizes specific sequence of DNA and cuts both the strands of DNA. For example, Eco RI recognizes the base sequences in DNA. REN may cut both the strands of DNA at the same position or at different positions. [When both the strands of DNA molecule are cut at the same position and no nucleotides are left unpaired, it results in blunt ends] [Symmetric ends] [When each strand of DNA molecule is cut at a different position, it results in sticky ends] [Asymmetric ends] e.g:- Eco RI, Hind III, Sal I, Bam 1.
Note: Restriction enzymes belong to a larger class of enzymes called nucleases. These are of two kinds : exonucleases and endonucleases. Exonucleases remove nucleotides from the ends of the DNA whereas, endonucleases make cuts at specific positions within the DNA.
(b) DNA Lygase:- The enzyme that joins the cut ends of DNA fragments is called DNA lygase. It is popularly called biomolecular stitchers [Molecular glues]. The enzyme joins either the blunt ends or the sticky ends of DNA.
4. Separation and isolation of DNA fragments:
The cutting of DNA by restriction endonucleases results in the fragments of DNA. These fragments can be separated by a technique known as gel electrophoresis. Since DNA fragments are negatively charged molecules they can be separated by forcing them to move towards the anode under an electric field through a medium/matrix. Nowadays the most commonly used matrix is agarose which is a natural polymer extracted from sea weeds. The DNA fragments separate (resolve) according to their size through sieving effect provided by the agarose gel. Hence, the smaller the fragment size, the farther it moves.
The separated DNA fragments can be visualised only after staining the DNA with a compound known as ethidium bromide followed by exposure to UV radiation (you cannot see pure DNA fragments in the visible light and without staining). You can see bright orange coloured bands of DNA in a ethidium bromide stained gel exposed to UV light. The separated bands of DNA are cut out from the agarose gel and extracted from the gel piece. This step is known as elution. The DNA fragments purified in this way are used in constructing recombinant DNA by joining them with cloning vectors.
5. Bioreactor:
It is a vessel designed to carry out biological reactions under controlled conditions. It is useful to regulate the contact between the catalyst and its substrate. Commercial bioreactors are generally used to produce the products on a large scale. It is made up of a stainless steel vessel or tank with some controlling units for PH, temperature and nutrient concentrations. It is also provided with pressure gauge inlets for nutrients and filtered air. It is also provided with a coil of pipe for letting in cold water or steam.
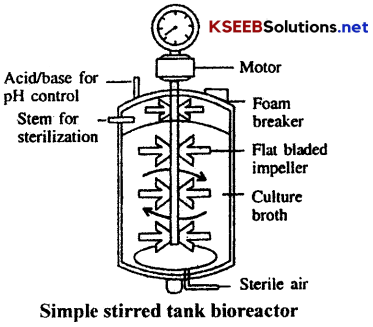
Uses of bioreactors:
- They are used to produce primary metabolites like ethanol, acetone, butanol, lysine, glutamic acid etc.
- They are used in the production of monoclonal antibodies, vaccines etc.
- They are used in tissue culture.
- They are used in shoot and root culture of some plants.
![]()
Process of Genetic Engineering:
(a) Isolation of genetic material (DNA): In recombinant DNA technology, it is essential to isolate DNA in pure form free from other macro molecules. Since DNA molecule is enclosed with the membrane in the cell. we have to break open the cell to release DNA along with other macromolecules like RNA, proteins, polysaccharides and lipids. This is carried out in bacterial cells, plant and animal cells with certain enzymes.
The other macro molecules can be removed by appropriate treatment with specific enzymes. Finally, the purified DNA molecules are precipiatated out after the addition of chilled ethanol and this can be seen as collection of fine threads in the suspension.
(b) Cutting of DNA at specific locations: The isolated purified DNA molecule is cut (cleaved) with the help of a suitable enzyme called restriction endonuclease, into segments with sticky ends.
(c) Gel electrophoresis: The cut DNA fragments are separated by gel electrophoresis using agarose gel. DNA is a negatively charged molecule, hence it moves towards the positive electrode (anode).
(d) Amplification of Gene of interest using PCR: Amplification of gene is ‘a process of making many copies of a gene’. It is achieved by using a technique called Polymerase Chain Reaction (PCR).
Requirements: 1
→ A DNA containing the desired segment to be amplified
→ Two nucleotide primers (about 20 base long) specific i.e. complementary to the two 3 – ends of the desired segment.
→ The four deoxynculeoside triphosphate i.e. TPP (Thymidine triphosphate) dCTP (Deoxycytidine triphosphate) dATP (deoxyadenosine triphosphate) and dGTP (deoxyguanosine triphosphate)
→ A thermostable DNA polymerase i.e., Taq DNA polymerase.
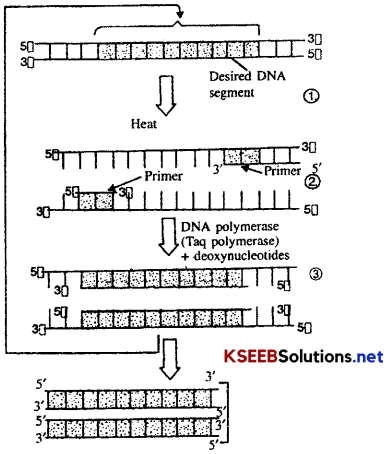
![]()
Procedure of PCR:
1. The DNA from the desired segment to be amplified, an excess of the two primer molecules, the four deoxyribose triphosphates and DNA polymerase are mixed together in a reaction mixture in a eppendorf tube with sufficient quantities of Mg++. The eppendorf tube is placed in the PCR unit and the following operations are performed sequentially.
2. Denaturation: The reaction mixture is first subjected to a temperature between 90-98°C (commonly 94°C). It results in the separation of two strands of DNA due to the breakage of hydrogen bonds. This is called denaturation. Each strand of DNA acts as a template strand for DNA synthesis. The duration of this step in the first cycle of PCR is usually 2 minutes at 94°C.
3. Annealing (anneal=join): The mixture is now cooled to a low temperature (40-60°C). During this step, two oligonucleotide primers, complementary to a region of DNA, anneal (hybridize) one to each 3 end of DNA strand. The duration of annealing step is usually one minute during the first as well as the subsequent cycles of PCR.
4. Primer extension: During this step, the enzyme taq DNA polymerase extend the primers using nucleotides and DNA templates. The two primers extent towards each other in order to get two new strands of DNA (at 5) end. The duration of primer extension is usually 2 minutes at 72°C. The amplified fragment, if required can now be used to ligate with vector for further cloning. The taq DNA polymerase remains active, during the high temperature induced denaturation of double stranded DNA.
(e) Insertion of recombinant DNA into the host: (competent host)
1. Electroporation: The bacterial cell is placed in a solution with cold CaCl2 solution followed by placing them at 42°C intermittently. This results in the development of pores in the cellmembrane. Now the recombinant plasmid migrates into the host cell and the bacterial cell gets transformed.
2. Microinjection : It is direct injection of a desired gene into the nucleus of an animal cell by micro syringe.
3. Biolistics : Here a suitable plant-cell is bombarded with high velocity microparticles (l-2nm) of gold or tungsten coated with DNA inorder to introduce DNA into the cell.
(f) Obtaining the Foreign Gene Product:
→ The transgene expresses itself in the form of protein (s) under appropriate conditions.
→ The product (s) can be extracted from the medium by employing a suitable procedure.
→ The transgenic cells may be cultured in the laboratory to obtain the transgene product on a small scale.
→ The transgenic cells can also be cultured/multiplied in a continuous culture system, in which the used medium is drained out from one side and fresh medium is added from the other side for the production of larger biomass and the desired product.
→ Bioreactors are used for processing large volumes of culture for obtaining the product of interest in sufficient quantities on a commercial scale.
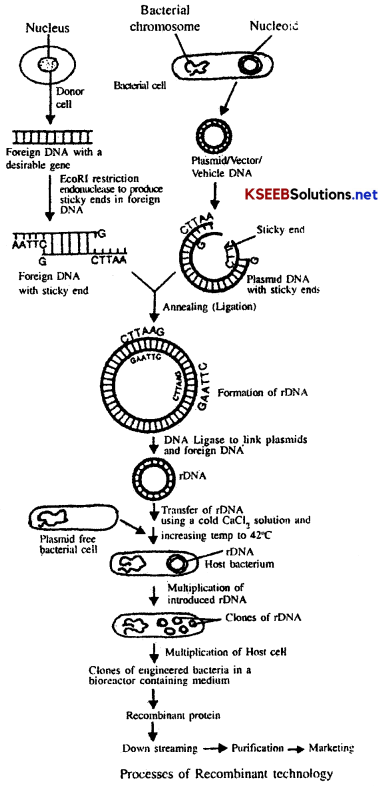
![]()
(g) Downstream Processing: The products formed in a bioreactor have to be subjected through a series of processes before they are ready for marketing as finished products. The various processes used for the recovery of useful products are collectively called downstream processing. The processes include separation and purification of the product, addition of suitable preservatives and a stringent quality control testing etc. Such formulation has to undergo strict trials as in the case of drugs. These quality control testing and clinical trials vary from product to product.